Checkpoints¶
TROVE provides an extended set of the so-called ‘checkpoints’, with the main functionality to provide the capability of restarting the calculations at any critical point. The checkpoints are TROVE files with extension .chk that contain the necessary information for the restart. Some of the files are in the ASCII format (i.e. ‘formated’), some are written in as machine readable, i.e ‘unformatted’, some - as direct access files. For instance, the transition between steps 1,2,3, … is driven by the checkpoint functionality. Moreover, since step 1 contains a number of critical sub-steps, TROVE checkpoints those sub-steps as well.
Apart from the restarting functionality, the .chk files are also used to store the eigenvectors (i.e. eigencoefficients). Although these objects are formally a the end result of the calculations, in same cases they provide support for some intermediate steps. For example, step 2 (transformation of matrix elements to the 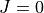 representation) used the eigenfunctions of the
representation) used the eigenfunctions of the 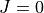 solution obtained at step 1 to create a basis set for step 3 (production of eigenfunctions for
solution obtained at step 1 to create a basis set for step 3 (production of eigenfunctions for  ).
).
The checkpoints control section is CHECK_POINT, e.g.:
CHECK_POINT
ascii
kinetic read
potential read
external none
basis_set save
contract save
matelem save split
extmatelem none split
eigenfunc save
END
with the main control keywords save, read and none and the additional keywords append, stitch, split etc. The functional checkpoint keywords include
kineticpotentialexternalbasis_setcontractmatelemextmatelemeigenfuncfit_poten
The order of these lines is unimportant. The CHECK_POINT section can appear anywhere in the input file.
List of checkpoints¶
kinetic.chk (
kinetic) contains the expansion coefficients of the KEO as controlled by theKinorderkeyword.The KEO is the first to be constructed and saved if
kineticis set tosaveand can be then used at any stage by switching toread. One opt out from saving the KEO (or any other checkpoint) by setting it tonone.potential.chk (
potential) is the second object created in TROVE as part of step 1.It contains expansion coefficient of PEF in terms of the TROVE vibrational coordinates. It can be saved, if generated from scratched and then read after a TROVE restart.
external.chk (
external) is used to store expansion coefficients of any other objects apart from KEO and PEF,dipole moment, polarizability, spin-rotation, a correction to the PEF used for the refinement of the PEF etc, which are called
ExternalorDipoleThe same functionality as above is applied to the External field.hamiltoian.chk:
This file contains the definition of the linearised coordinates (A and B matrices, see the TROVE paper [TROVE]), i.e. for
COORDS linearin the case of the linearised coordinate. It is saved and read as part of thekineticswitch. hamiltoian.chk is not produced for the curvilinear coordinate typeCOORDS local.kinetic.chk, potential.chk and external.chk have the ASCII format and can be used to plot the corresponding fields or their individual components.
numerov_bset.chk and prim_bset.chk (
basis_set).These two unformatted files containing information on the 1D primitive basis sets and are produced after the objects described above. The same rules are applied for saving, reading or doing nothing (
none).contr_descr.chk, contr_quanta.chk, contr_vectors.chk (
contract) are three files to store the information on the symmetry adapted basis functions for individual basis sub-sets.contr_vectors.chk is an unformatted file containing eigne-coefficients.
contr_quanta.chk is a formatted (ASCII) file with the information on the basis set bookkeeping - mapping of the multimode quantum numbers into a 1D array.
contr_descr.chk is a formatted file containing descriptions of the basis sets:
individual energies of the basis functions from different sub-sets together with their classifications, symmetries, TROVE quantum number, IDs, largest expansion coefficients used in their assignment as well as a placeholder for the spectroscopic quantum numbers. This file can be edited in order to include these spectroscopic quantum numbers.
contr_matelem.chk (
matelem) contains vibrational matrix elements of the different pars of the Hamiltonian operator (KEO and PEF)on the basis set produced at the
contractstep, i.e. contracted symmetry adapted basis functions.matelem1.chk, matelem2.chk … matelem12.chk represent a
splitversion of contr_matelem.chk, which contain the rotational and Coriolis parts of the KEO,while contr_matelem.chk contains the matrix elements of the pure vibrational part of the Hamiltonian operator, i.e. vibrational KEO + PEF. The
splitoption allows one to compute all these 13 parts ofmatelemsseparately, which is especially useful for larger molecules. Historically, thematelemstep the main bottleneck in the TROVE pipeline in the case of large molecules and thesplitfeature allows one to at least partly mitigate this issue. An example of thesplitoption isTo process all 13 parts of the Hamiltonian:
contract save split
To process individual parts of the Hamiltonian (from 0 to 6):
contract save split 0 6 here 0 stands for the pure vibrational part of the Hamiltonian operator.
To process a single part of the Hamiltonian (from 11 to 11):
contract save split 11 11
contr_extfield.chk
extmatelemcontains all (vibrational) matrix elements of the external field.extmatelemstep is not a compulsory step in the TROVE pipeline. It is invoked when keyword it is set tosave.extmatelem1.chk, extmatelem2.chk, extmatelem3.chk …. are the
splitanalogy of contr_extfield.chk,where different components are written into separate extmatelem chk-files.
Examples of the split option include :
extmatelem save split 1 1
:
extmatelem save split
eigen_descr*chk, eigen_vector*chk and eigen_quant*chk (
eigenfunc) contain the eigencoefficients and their descriptions.eigen_descr
 _
_ .chk contain the eigenvalues (energy term values in cm-1):
.chk contain the eigenvalues (energy term values in cm-1):
state IDs, symmetries, TROVE quantum numbers, largest coefficients as well as a placeholder for the spectroscopic quantum numbers. These files formatted (ASCII) and can be used for the analysis or postprocessing (e.g. construction of line lists). Here
 is the rotational angular momentum and
is the rotational angular momentum and  is the symmetry (irrep), i.e. there is a description for each J/symmetry. For example, eigen_descr0_2.chk is a checkpoint file with the description of the eigenstates and their eigenvalues for
is the symmetry (irrep), i.e. there is a description for each J/symmetry. For example, eigen_descr0_2.chk is a checkpoint file with the description of the eigenstates and their eigenvalues for 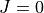 and
and  .
.eigen_vectors
 _
_ .chk contain the eigencoefficients written in direct unformatted form.
.chk contain the eigencoefficients written in direct unformatted form.For each eigen_descr*chk there is an eigen_vector*chk file.
eigen_quant
 .chk contain the bookkeeping information for the basis sets used:
.chk contain the bookkeeping information for the basis sets used:the mapping between the multidimensional, multimode description of the product-form basis functions to a 1D basis set index.
j0_matelem.chk (
matelem) is the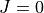 representation of contr_matelem.chk generated at step 2.
representation of contr_matelem.chk generated at step 2.In order to switch to step 2 and thus distinguish from step 1, the following changes to the step 1 input file should be made:
In the
contractedsection, setmodel J=0
In the
check_pointsection set.... contract save matelem convert eigenfunc read ....
j0_matelem1.chk, j0_matelem2.chk … j0_matelem12.chk are the
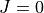 representation of matelem1.chk, matelem2.chk … matelem12.chk, respectively, in the
representation of matelem1.chk, matelem2.chk … matelem12.chk, respectively, in the splitform.These files are generated as part of step 2, which can be accomplished by simply setting
step 2in theControlsection:control step 2 end
Alternatively, the changes described above to produce j0_matelem.chk should be introduced, with only one difference of including the
splitsub-option:.... contract save matelem convert split eigenfunc read ....
In the
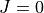 representation, the zero-term, pure vibrational j0_matelem0.chk is not produced. This is because this part is diagonal on the
representation, the zero-term, pure vibrational j0_matelem0.chk is not produced. This is because this part is diagonal on the 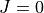 basis, with the corresponding energies on the diagonal.
basis, with the corresponding energies on the diagonal.fitpot_ma
 _
_ _:math:i.chk (
_:math:i.chk (fit_poten) are checkpoint files to store matrix elements of the fitting part of the potential, with representing the vibrational contribution from a specific potential parameter
representing the vibrational contribution from a specific potential parameter  to be refined.
to be refined.To produce and store the fitpot*.chk checkpoints, the following line should be added to the
check_pointsection:..... fit_poten save split .....
and to use them
..... fit_poten read split .....
Similar to the usage of other
splitobjects, it is possible to request only specific terms in the potential, e.g..... fit_poten save split 32 45 ....
will compute the
fit_potenmatrix elements for .
.The matrix elements in fitpot*.chk are used for the refinement of the PEF, which is controlled by the section
FITTING, see Chapter “Refine”. This section contains keywords for selection of fitpot*.chk, namelyJ-LISTandsymmetriesspecifying the values of and symmetries
and symmetries  , respectively (both are integer) to be processed. For example:
, respectively (both are integer) to be processed. For example:FITTING J-LIST 0 1 2 Symmetries 2 3 5 6 .....
will process the fit (
fit_potenmatrix elements) for :math`J=0,1,2` and .
.
“Fingerprints”¶
In order to help prevent using wrong checkpoints from different project, the descr checkpoint files (contr_descr.chk, eigen_descr*_*.chk and j0eigen_descr_*.chk) contain a “fingerprint” section at the very beginning of these formatted (ASCII) files.
Here is an example of the top part of a file eigen_descr0_3.chk from an H2S project:
Start Fingerprints
0 3 3 4 38000.0 38000.0 <= PTorder, Nmodes, Natoms, Npolyads, enercut
0.01000000 0.01000000 0.01000000 <= dstep
31.97207070 1.00782505 1.00782505 <= masses
1.33590070 1.33590070 1.61034345 <= equilibrium
LINEAR R-RHO C2V(M) XY2
BASIS: i type coord_kinet coord_poten model dim species class range dvrpoints res_coeffs npoints borders periodic period
0 <- Jrot, rotational angular momentum
0 JKTAU xxxxxx xxxxxx 1000 1D 0 0 0 0 0.0 0 0.000 0.000 F 0 xxxxxx 0 F F T
1 NUMEROV LINEAR MORSE 1000 1D 1 1 0 4 1.0 600 -0.500 1.400 F 0 NUMEROV-PO 5 F F T
2 NUMEROV LINEAR MORSE 1000 1D 1 1 0 4 1.0 600 -0.500 1.400 F 0 NUMEROV-PO 5 F F T
3 NUMEROV LINEAR LINEAR 1000 1D 2 2 0 4 1.0 4000 0.070 2.618 F 0 NUMEROV-PO 5 F F T
End Fingerprints
Start Quantum numbers and energies
4 <== Contracted polyad number
.........
.........
End Quantum numbers and energies
This information is automatically degenerated using some key parameters from the input project in question, such as atomic masses, number of atoms, maximal polyad, frame, coordinates type, symmetry and a detailed description of the basis set. When such a checkpoint is read by TROVE, the fingerprint is compared against the corresponding parameters of the current project. TROVE will stop and report in case of any differences found.
These descr files end with an “End …” section, which is also used to check if all the information required has been read correctly.
Similar preventive check are used in unformatted files as well, which also start with a section “Start … “ and end with a section “End…”. These are useful to catch files that are too short or long for a given project.
Structure of the description checkpoints¶
contr_descr.chk¶
contr_descr.chk contains the energies and quantum numbers of the contracted basis states. It has the following structure
1. “Fingerprint” section, for example
Start Fingerprints 0 3 3 4 38000.0 38000.0 <= PTorder, Nmodes, Natoms, Npolyads, enercut 0.01000000 0.01000000 0.01000000 <= dstep 31.97207070 1.00782505 1.00782505 <= masses 1.33590070 1.33590070 1.61034345 <= equilbrium LINEAR R-RHO C2V(M) XY2 BASIS: i type coord_kinet coord_poten model dim species class range dvrpoints res_coeffs npoints borders periodic period 0 <- Jrot, rotational angular momentum 0 JKTAU xxxxxx xxxxxx 1000 1D 0 0 0 0 0.0 0 0.000 0.000 F 0 xxxxxx 0 F F T 1 NUMEROV LINEAR MORSE 1000 1D 1 1 0 4 1.0 600 -0.500 1.400 F 0 NUMEROV-PO 5 F F T 2 NUMEROV LINEAR MORSE 1000 1D 1 1 0 4 1.0 600 -0.500 1.400 F 0 NUMEROV-PO 5 F F T 3 NUMEROV LINEAR LINEAR 1000 1D 2 2 0 4 1.0 4000 0.070 2.618 F 0 NUMEROV-PO 5 F F T End Fingerprints2. Sub-class 1
Class # 1 15 15 <- number of roots and dimension of basis 1 1 1 1 3631.457962557468 0 0 0 0 0 0 0 0 0.99650855 2 1 2 1 6237.793846833698 0 1 0 0 0 1 0 0 -0.70449312 3 4 3 1 6920.808287732269 0 0 1 0 0 1 0 0 -0.69933488 4 1 4 1 8799.405574657818 0 1 1 0 0 2 0 0 -0.66971512 5 4 5 1 9424.198345933781 0 0 2 0 0 2 0 0 -0.69495690 6 1 6 1 10131.498717237881 0 1 1 0 0 2 0 0 0.72224834 7 1 7 1 11335.796189988198 0 2 1 0 0 3 0 0 -0.57998640 8 4 8 1 11965.585881042278 0 0 3 0 0 3 0 0 0.625724312. Sub-class 2
Class # 2 5 5 <- number of roots and dimension of basis 16 1 1 1 3676.343781669158 0 0 0 0 0 0 0 0 0.99914576 17 1 2 1 4863.062238559608 0 0 0 1 0 0 0 1 -0.99741510 18 1 3 1 6044.794034517188 0 0 0 2 0 0 0 2 -0.99558857 19 1 4 1 7223.358797674692 0 0 0 3 0 0 0 3 -0.99426327 20 1 5 1 8404.773619941834 0 0 0 4 0 0 0 4 0.99669840
etc. Here the TROVE sub-class stands for a symmetry independent sub-set.
The structure of the columns is as follows:
-- ------ ---- ---- ------------------ --- ---- --- ---- --- ---- --- ------ -------------
1 2 3 4 5 6 7 8 9 10 11 12 13 14
-- ------ ---- ---- ------------------ --- ---- --- ---- --- ---- --- ------ -------------
n Sym m deg Energy (cm-1) k v1 v2 v3 K n1 n2 n3 C_i
-- ------ ---- ---- ------------------ --- ---- --- ---- --- ---- --- ------ -------------
1 1 1 1 3631.457962557468 0 0 0 0 0 0 0 0 0.99650855
2 1 2 1 6237.793846833698 0 1 0 0 0 1 0 0 -0.70449312
3 4 3 1 6920.808287732269 0 0 1 0 0 1 0 0 -0.69933488
4 1 4 1 8799.405574657818 0 1 1 0 0 2 0 0 -0.66971512
5 4 5 1 9424.198345933781 0 0 2 0 0 2 0 0 -0.69495690
where
Col 1:
nis the counting number of the state including degeneracies;Col 2:
Symis the irrep of this state;Col 3:
mis the running number of the energy, excluding degeneracies.Col 4:
degis a degenerate component (e.g.and

2for a 2D irrep of C3v).Col 5:
Energyterm value of a state from the given sub-class.Col 6:
kis a rotational index (typically it is zero in contr_descr.chk).Cols 7-9:
v1,v2,v3are the TROVE (local mode) vibrational quantum numbers.Col 10-13:
K, n1, n2, n3are placeholder for the user-defined quantum numbers to be propagated to the final rovibrational eigenstates.Col 14:
C_iis the corresponding largest coefficient used in the assignment.
eigen_descr*_*.chk¶
The eigenvector igen_descr _
_ .chk are used to store information on the ro-vibrational states. Here
.chk are used to store information on the ro-vibrational states. Here  and
and  are the rotational angular momentum and the symmetry, respectively.
are the rotational angular momentum and the symmetry, respectively.
1. “Fingerprint” section, for example:
Start Fingerprints 0 3 3 4 38000.0 38000.0 <= PTorder, Nmodes, Natoms, Npolyads, enercut 0.01000000 0.01000000 0.01000000 <= dstep 31.97207070 1.00782505 1.00782505 <= masses 1.33590070 1.33590070 1.61034345 <= equilbrium LINEAR R-RHO C2V(M) XY2 BASIS: i type coord_kinet coord_poten model dim species class range dvrpoints res_coeffs npoints borders periodic period 0 <- Jrot, rotational angular momentum 0 JKTAU xxxxxx xxxxxx 1000 1D 0 0 0 0 0.0 0 0.000 0.000 F 0 xxxxxx 0 F F T 1 NUMEROV LINEAR MORSE 1000 1D 1 1 0 4 1.0 600 -0.500 1.400 F 0 NUMEROV-PO 5 F F T 2 NUMEROV LINEAR MORSE 1000 1D 1 1 0 4 1.0 600 -0.500 1.400 F 0 NUMEROV-PO 5 F F T 3 NUMEROV LINEAR LINEAR 1000 1D 2 2 0 4 1.0 4000 0.070 2.618 F 0 NUMEROV-PO 5 F F T End Fingerprints 2. "Quantum numbers and energies" section, Energy term values and description of the ro-vibrational states: :: Start Quantum numbers and energies 4 <== Contracted polyad number Class # 1 22 22 <- number of roots and dimension of basis 1 1 1 1 3627.658022299023 0 0 0 0 1 1 1 1 0 0 0 0 0.99958266 1 1 2 1 2 1 4800.325667905056 0 0 0 1 2 1 1 1 0 0 0 1 -0.99819947 1 2 3 1 3 1 5962.955540973960 0 0 0 2 3 1 1 1 0 0 0 2 0.99055229 1 3 4 1 4 1 6236.371962631670 0 1 0 0 6 1 1 1 0 1 0 0 0.99283316 2 1 5 1 5 1 7130.700436920808 0 0 0 3 4 1 1 1 0 0 0 3 0.97608301 1 4 6 1 6 1 7393.117966742439 0 1 0 1 7 1 1 1 0 1 0 1 0.97535336 2 2 ...... ...... End Quantum numbers and energies
The meaning of the state columns is as follows:
---- ------- ------- ------ -------------------- ----- --- --- ----- ----- --- --- ----- ----- --- --- ------ ---------------- ----- ------
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20
---- ------- ------- ------ -------------------- ----- --- --- ----- ----- --- --- ----- ----- --- --- ---- - ---------------- ----- ------
n Sym m deg Energy (cm-1) k v1 v2 v3 imax s0 s1 s2 K n1 n2 n3 C_i ivib1 ivib2
---- ------- ------- ------ -------------------- ----- --- --- ----- ----- --- --- ----- ----- --- --- ---- ------------------ ----- ------
1 1 1 1 3627.658022299023 0 0 0 0 1 1 1 1 0 0 0 0 0.99958266 1 1
2 1 2 1 4800.325667905056 0 0 0 1 2 1 1 1 0 0 0 1 -0.99819947 1 2
3 1 3 1 5962.955540973960 0 0 0 2 3 1 1 1 0 0 0 2 0.99055229 1 3
.................
Here
Col 1:
nis the counting number of the state including degeneracies;Col 2:
Symis the irrep of this state;Col 3:
mis the running number of the energy, excluding degeneracies.Col 4:
degis a degenerate component (e.g.
1and
2for a 2D irrep of C3v).Col 5:
Energyterm value (cm-1).Col 6:
kis a rotational index.Cols 7-9:
v1,v2,v3are the TROVE (local mode) vibrational quantum numbers.Col 10:
imaxis the position index of the largest coefficient in the basis vector.Col 11:
s0is the symmetry of the rotational basis set contribution for the term with the largest eigen-coefficient.Col 12-13:
s1ands2are symmetries of the vibrational contribution for each sub-space.Col 14-17:
K, n1, n2, n3are placeholder for the user-defined quantum numbers to be propagated to the final rovibrational eigenstates.Col 18:
C_iis the corresponding largest coefficient used in the assignment.Col 19-20:
ivib1,ivib2are counting indices of sub-classes in the representation of direct products of the symmetry adapted ‘contracted’ basis set.
j0eigen_descr*_*.chk¶
The eigenvector j0eigen_descr _
_ .chk are used to store information on the ro-vibrational states in the
.chk are used to store information on the ro-vibrational states in the 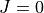 representation. Here
representation. Here  and
and  are the rotational angular momentum and the symmetry, respectively. The general structure of the ‘j0eigen_descr’ checkpoint files is the same as for ‘eigen_descr, a fingerprint section is followed by the description section with “Quantum numbers and energies”. The latter in
are the rotational angular momentum and the symmetry, respectively. The general structure of the ‘j0eigen_descr’ checkpoint files is the same as for ‘eigen_descr, a fingerprint section is followed by the description section with “Quantum numbers and energies”. The latter in j0eigen_descr*_*.chk section is only slightly different from that from eigen_descr*_*.chk. The 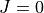 representation contains a single one-sub class with full vibrational (
representation contains a single one-sub class with full vibrational (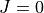 ) basis functions.
) basis functions.